Single-cell transcriptomic analyses reveal distinct immune cell contributions to epithelial barrier dysfunction in checkpoint inhibitor colitis
Molly Fisher Thomas1-5*†, Kamil Slowikowski1-4*†, Kasidet Manakongtreecheep1-3, Pritha Sen1-4,6, Nandini Samanta1-3, Jessica Tantivit1-3, Mazen Nasrallah1-4,7, Leyre Zubiri2,4, Neal P. Smith1-3, Alice Tirard1-3, Swetha Ramesh1-3, Benjamin Y. Arnold1-3, Linda T. Nieman2, Jon athan H. Chen2-4,8, Thomas Eisenhaure3, Karin Pelka2-4, Yuhui Song2, Katherine H. Xu2, Vjola Jorgji2,8, Christopher J. Pinto2,9, Tatyana Sharova9, Rachel Glasser5, PuiYee Chan2,4, Ryan J. Sullivan2,4, Hamed Khalili3-5, Dejan Juric2-4, Genevieve M. Boland2,4,10, Michael Dougan4-5, Nir Hacohen2-4, Bo Li1,3,11, Kerry L. Reynolds2,4‡, and Alexandra-Chloé Villani1-4,7‡†
Author affiliations
- 1 - Center for Immunology and Inflammatory Diseases, Department of Medicine, Massachusetts General Hospital, Boston, MA, USA
- 2 - Massachusetts General Hospital, Cancer Center, Boston, MA, USA
- 3 - Broad Institute of Massachusetts Institute of Technology and Harvard, Cambridge, MA, USA
- 4 - Harvard Medical School, Boston, MA, USA
- 5 - Division of Gastroenterology, Department of Medicine, Massachusetts General Hospital, Boston, Massachusetts, USA
- 6 - Division of Infectious Disease, Department of Medicine, Massachusetts General Hospital, Boston, MA, USA
- 7 - Division of Rheumatology, North Shore Physicians Group, Department of Medicine, Mass General Brigham Healthcare Center, Lynn, MA, USA
- 8 - Department of Pathology, Massachusetts General Hospital, Boston, MA, USA
- 9 - Clinical Research Center, Massachusetts General Hospital, Boston, MA, USA
- 10 - Department of Surgery, Massachusetts General Hospital, Boston, MA, USA
- 11 - Current address: Genentech, 1 DNA Way, South San Francisco, CA, 94080
* These authors contributed equally: Molly Fisher Thomas, Kamil Slowikowski
‡These authors contributed equally: Kerry Reynolds, Alexnandra-Chloé Villani
† Correspondence to: mfthomas@partners.org; kslowikowski@mgh.harvard.edu; avillani@mgh.harvard.edu
Nature Medicine 2024. doi: 10.1038/s41591-024-02895-x
Abstract
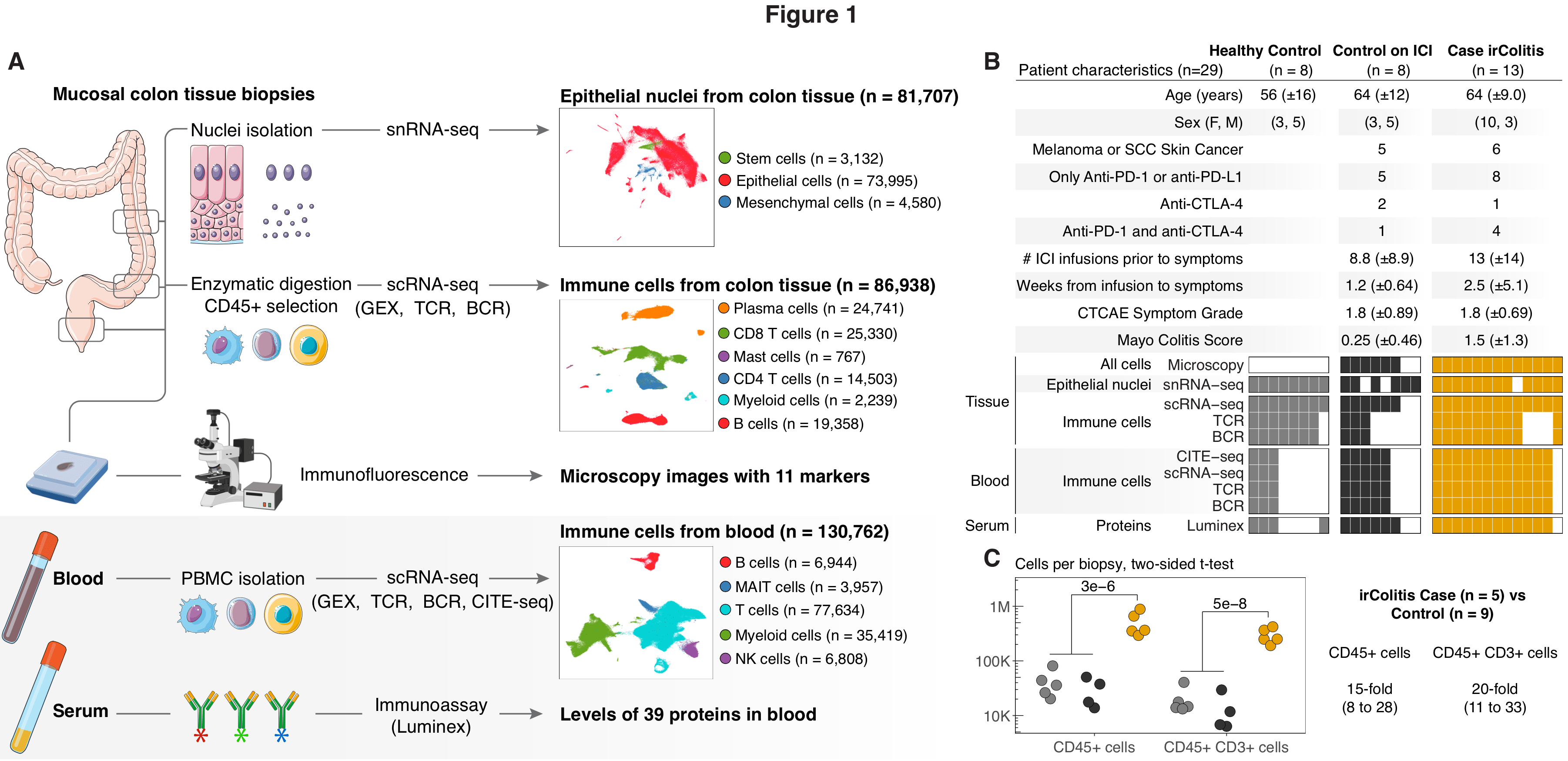
Therapeutic immune checkpoint blockade has revolutionized oncology, but treatments are limited by immune-related adverse events, including checkpoint inhibitor colitis (irColitis). To define molecular drivers of irColitis, we profiled ~300,000 cells from the colon mucosa and blood of 29 patients and controls. Patients with irColitis showed expanded mucosal Tregs, ITGAEHi CD8 tissue-resident memory T cells expressing CXCL13 and Th17 gene programs, and circulating ITGB2Hi CD8 T cells recruited by ICAM and CXCR3 ligands. Cytotoxic GNLYHi CD4 T cells, ITGB2Hi CD8 T cells from circulation, and endothelial cells associated with hypoxia gene programs were further expanded in colitis associated with anti-PD-1/CTLA-4 compared to anti-PD-1 therapy. Luminal epithelial cells in irColitis patients expressed PD-L1 and interferon-induced signatures associated with apoptosis, increased cell turnover, and malabsorption. Together, we highlight novel roles for circulating T cells and epithelial-immune crosstalk critical to PD-1/CTLA-4-dependent tolerance and barrier function and nominate novel irColitis therapeutic targets.
🔬 View the data
On this website, we provide interactive data browsers to view all of the transcriptomics data for each of the manually curated cell clusters.
Cell Clusters
Metadata variables and gene expression in two-dimensional embeddings.
Tissue immune cells:
- CD8 T cells, CD4 T cells, Myeloid cells, B cells
Tissue epithelial and mesenchymal nuclei:
- 23 types of epithelial and mesenchymal nuclei
- CD8 T cells, CD4 T cells, Myeloid cells, B cells
Gene Contrasts
Differential expression statistics for all genes:
- 3 contrasts
-
- irColitis Case vs Control
- Case PD1/CTLA4 vs Case PD1
- Control PD1 vs Control None
- 9 major cell lineages
- 105 cell clusters
📝 Cite our work
Thomas MF, Slowikowski K, Manakongtreecheep K, Sen P, Samanta N, Tantivit J, et al. Single-cell transcriptomic analyses reveal distinct immune cell contributions to epithelial barrier dysfunction in checkpoint inhibitor colitis. Nature Medicine. 2024; 1–14. doi:10.1038/s41591-024-02895-x
BibTeX
@ARTICLE{Thomas2024-lk,
title = "{Single-cell transcriptomic analyses reveal distinct immune cell
contributions to epithelial barrier dysfunction in checkpoint
inhibitor colitis}",
author = "Thomas, Molly Fisher and Slowikowski, Kamil and
Manakongtreecheep, Kasidet and Sen, Pritha and Samanta, Nandini
and Tantivit, Jessica and Nasrallah, Mazen and Zubiri, Leyre and
Smith, Neal P and Tirard, Alice and Ramesh, Swetha and Arnold,
Benjamin Y and Nieman, Linda T and Chen, Jonathan H and
Eisenhaure, Thomas and Pelka, Karin and Song, Yuhui and Xu,
Katherine H and Jorgji, Vjola and Pinto, Christopher J and
Sharova, Tatyana and Glasser, Rachel and Chan, Puiyee and
Sullivan, Ryan J and Khalili, Hamed and Juric, Dejan and Boland,
Genevieve M and Dougan, Michael and Hacohen, Nir and Li, Bo and
Reynolds, Kerry L and Villani, Alexandra-Chloé",
journal = "Nature Medicine",
publisher = "Nature Publishing Group",
pages = "1--14",
abstract = "Immune checkpoint inhibitor (ICI) therapy has revolutionized
oncology, but treatments are limited by immune-related adverse
events, including checkpoint inhibitor colitis (irColitis).
Little is understood about the pathogenic mechanisms driving
irColitis, which does not readily occur in model organisms, such
as mice. To define molecular drivers of irColitis, we used
single-cell multi-omics to profile approximately 300,000 cells
from the colon mucosa and blood of 13 patients with cancer who
developed irColitis (nine on anti-PD-1 or anti-CTLA-4 monotherapy
and four on dual ICI therapy; most patients had skin or lung
cancer), eight controls on ICI therapy and eight healthy
controls. Patients with irColitis showed expanded mucosal Tregs,
ITGAEHi CD8 tissue-resident memory T cells expressing CXCL13 and
Th17 gene programs and recirculating ITGB2Hi CD8 T cells.
Cytotoxic GNLYHi CD4 T cells, recirculating ITGB2Hi CD8 T cells
and endothelial cells expressing hypoxia gene programs were
further expanded in colitis associated with anti-PD-1/CTLA-4
therapy compared to anti-PD-1 therapy. Luminal epithelial cells
in patients with irColitis expressed PCSK9, PD-L1 and
interferon-induced signatures associated with apoptosis,
increased cell turnover and malabsorption. Together, these data
suggest roles for circulating T cells and epithelial–immune
crosstalk critical to PD-1/CTLA-4-dependent tolerance and barrier
function and identify potential therapeutic targets for
irColitis. Single-cell multi-omic analysis of 300,000 cells from
29 patients representing peripheral immune cells and colon
mucosal immune, epithelial and mesenchymal cells reveals
crosstalk between circulating and tissue-resident immune cells
with epithelial cells in checkpoint inhibitor colitis and
identifies potential therapeutic targets.",
month = may,
year = 2024,
doi = "10.1038/s41591-024-02895-x",
issn = "1546-170X,1546-170X",
language = "en"
}
💾 Download the data
Analysis result files are available at Zenodo: 10.5281/zenodo.8088435
Single-cell expression data files are available at NCBI GEO: GSE206301
Sequencing reads are available at dbGAP: phs003418.v1.p1
💻 Read the source code
The code for our analysis and for this website is available at GitHub: github.com/villani-lab/ircolitis
Learn more about the data files we shared: github.com/villani-lab/ircolitis/analysis/output/README.md
✉ Contact us
Please contact us with any questions or comments.
The data presented here comes from the laboratory of Dr. Alexandra-Chloé Villani.
📰 In the news
- Nature Medicine Research Briefing: Studying paired patient tissue and blood enables insights into immunotherapy toxicity
- Research Spotlight: A Better Understanding of Immune Checkpoint Inhibitor-Related Colitis
- LinkedIn post by Broad Institute of MIT and Harvard
- Twitter post by MGH Research Institute